Cryo-EM for Subcellular Organelles
The rapid development of cryogenic electron microscopy (cryo-EM) in the structural determination field suggests that the structural analysis of molecular components in entire cells and subcellular organelles is about to enter a new era. The electron tomography technology can be used to study the internal structure of subcellular organelles with a certain thickness in the natural state. Our cryo-EM platform is constantly learning to identify sub-organelles more accurately, and we continue to make breakthroughs in cytoskeleton, nuclear pore complex, and other sub-organelles.
Cryo-Electron Tomography (Cryo-ET), one of the techniques of Cryo-EM, is used to produce three-dimensional (3D) pictures from a series of tilted Cryo-EM images. Cryo-ET has gained more and more attention in the field of Cryo-EM in the recent years, which can be applied to study the internal organisms, typically for many structures inside cells, for example, subcellular organelles. Organelle is a specialized subunit within a cell, which contains cytoskeleton, mitochondria, endoplasmic reticulum, flagellum and so on. Cryo-EM allows to study the high-resolution structures of macromolecules, and Cryo-ET provides the information about cellular processes. With the combination of 3D imaging and specimen preparation that preserves the integrity of the cell structure, Cryo-ET can observe the internal structure of cell with high resolution.
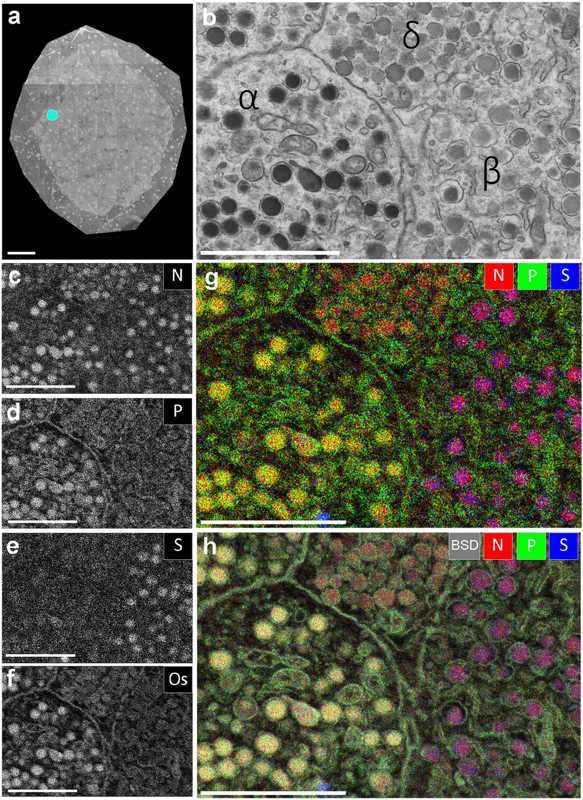 Figure 1. EDX defines cell types and subcellular structures and organelles in EM images (Scotuzzi M, et al. 2017)
Figure 1. EDX defines cell types and subcellular structures and organelles in EM images (Scotuzzi M, et al. 2017)
The following is a general process of cryo-EM analysis on cell sub-organelles we provide for you:
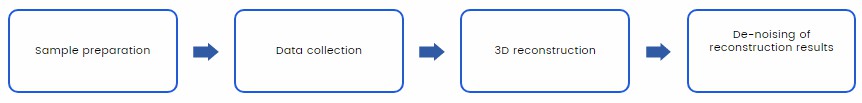
At Creative Biostructure, we have the most advanced Electron Microscopes equipment and software. In addition, Creative Biostructure has a professional team for experimental design of subcellular organelles separation and Cryo-ET data analysis.
The advantages of our service include:
- The most advanced direct-electron detectors and image processing software;
- Allow near-atomic resolution structures to be calculated without crystallization and only need a small amount of purified samples;
- Provide the unique possibility to image organelle in their native environment or in situ;
- Visualize the interaction of proteins and cells, capture transient and dynamic processes;
- Bring a hope for further understanding of life mechanisms and mysteries.
Our cryo-EM team is very happy to provide cryo-EM structure analysis services for subcellular organelle samples. If you are interested in our services, please feel free to contact us. We are looking forward to cooperating with you.
Ordering Process
- Scotuzzi M, et al. Multi-color electron microscopy by element-guided identification of cells, organelles and molecules. Scientific Reports. 2017. 7(1): 1-8.
