Randomized Mutagenesis Libraries
Supported by our advanced protein engineering platform established for years, Creative Biostructure has developed various approaches for mutagenesis libraries construction, including gene and protein variants. Our professional team provides custom randomized mutagenesis libraries construction services with a complexity of up to 109.
Protein mutant libraries are composed of thousands of different protein variants and each variant in the library contains different mutations. In other words, they are collections of DNA oligonucleotides (oligoes) which are generally used for creating families of variant genes and proteins for functional studies. Protein mutagenesis libraries have been generally used to optimize protein functions, by recognizing and modifying crucial residues so as to alter the protein structurally and functionally. It is an important approach of protein engineering to generate newfangled proteins, moreover, by using this approach the solubility, stability and expression levels of proteins greatly improved.
Randomized mutant libraries are produced by de novo synthesis of a gene, during the process, either nucleotides or codons are coalescent into the DNA sequence at specific positions. Finally, a mixture of double-stranded DNA molecules is obtained, which stand for variants of the original gene. Therefore, individual clones enable to be founded by incorporating the variants into an expression vector, and the encoded protein variants can be expressed.
Creative Biostructure offers randomized mutagenesis libraries construction services with library complicacy up to 109 delivered as dsDNA, clones with plasmid vector your chose, or transformants in glycerol stocks.
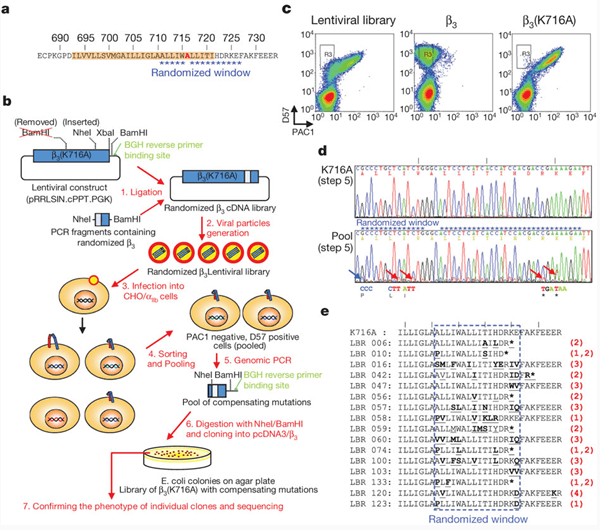 Figure 1: Directed evolution of the β3 integrin to identify mutations that complement the activating effect of snorkelling lysine mutation.
Figure 1: Directed evolution of the β3 integrin to identify mutations that complement the activating effect of snorkelling lysine mutation.
- Strategies for Randomized Mutagenesis Libraries Construction
In vitro library synthesis and degenerated oligonucleotide technology are able to introduce random mutations on a steerable level with maximum elasticity, allowing highly precise randomization within oligonucleotides. Based on the progressive degenerated oligonucleotide techniques, Creative Biostructure can perform any form of randomization of the full-length gene in a synthetic DNA fragment.
- Approaches for Randomized Mutagenesis Libraries Construction
Creative Biostructure has developed several methods to create random mutant libraries, such as error-prone PCR (epPCR), rolling circle error-prone PCR, sequence saturation mutagenesis (SeSaM), mutator strains, temporary mutator strains, insertion mutagenesis, ethyl methanesulfonate and nitrous acid. Our protein randomized mutagenesis libraries construction services contain gene synthesis, mutagenesis, sub-cloning, protein expression, and finally, protein purification.
Creative Biostructure provides various mutagenesis libraries construction services. Please feel free to contact us to discuss your project!
Ordering Process
References:
Drummond D A, Iverson B L, Georgiou G, et al. Why high-error-rate random mutagenesis libraries are enriched in functional and improved proteins[J]. Journal of molecular biology, 2005, 350(4): 806-816.
Abou-Nader M, Benedik M J. Rapid generation of random mutant libraries[J]. Bioengineered bugs, 2010, 1(5): 337-340.
Cadwell R C, Joyce G F. Randomization of genes by PCR mutagenesis[J]. Genome research, 1992, 2(1): 28-33.
Pai J C, Entzminger K C, Maynard J A. Restriction enzyme-free construction of random gene mutagenesis libraries in Escherichia coli[J]. Analytical biochemistry, 2012, 421(2): 640-648.
